Peptigram @ Bioware
Welcome to Peptigram, a user friendly web application for peptidomics data visualization.
Peptigram is freely available for academic use. For other use please contact us.
What kind of visualisation does Peptigram produce?
Peptigram produces two kinds of visualisation for every precursor protein:
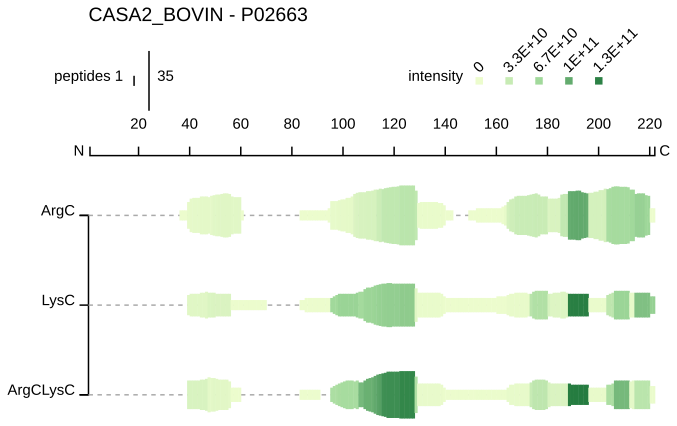
- peptide profile, peptide coverage visualisation with intensity values for every sample.
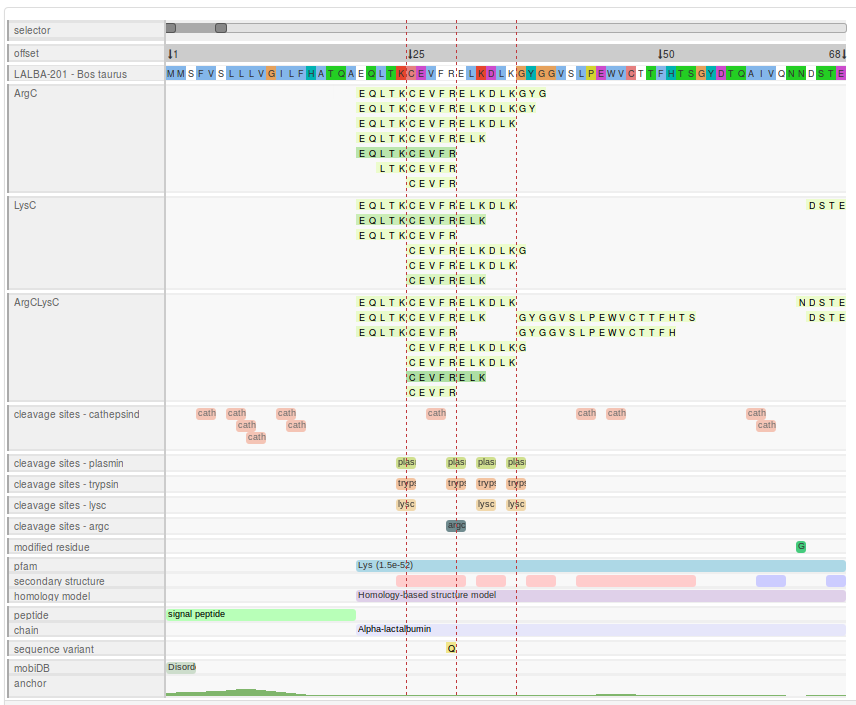
- peptide alignment map, peptide sequences for every samples are mapped to the precursor protein sequence along with other information about the protein and proteolytic cleavage sites.
Every visualisation is interactive, customizable and can be downloaded.
How to use Peptigram?
- create a new job and upload your data
- create peptide profiles
- select a precursor protein to create a peptide alignment map
- modify it as you wish
What kind of data does Peptigram use?
Your data
Peptigram only requires you to submit a CSV file the information relating to the peptides you want to visualize (peptide sequences, UniProt IDs, start, end, intensities). Multiple samples can be compared, you just have to add an intensity column to your CSV file, Peptigram will do the rest. Full specifications and more information are available in the documentation.
External data
Peptigram uses ProViz to retrieve data from public protein data repositories (e.g. UniProt and GeneTree). Peptigram also uses cleavage sites from PeptideCutter (ExPASy) and from Enzyme Predictor.
Download test dataWho makes Peptigram?
Peptigram is currently developed by Jean Manguy from the Shields lab, UCD Conway Institute of Biomolecular and Biomedical Research; funded by Food for Health Ireland and by the Irish Research Council.
If you use Peptigram please cite the following article.
Disclaimer
Peptigram is freely available for academic research. For other usages, please contact us.
Peptigram works best with most standard compliant web browsers. We recommend the use of Firefox, Chrome, Opera or Safari. Although we succesfully tested most of the features on Microsoft Edge, we do advise against the use of this browser. Internet Explorer is outdated and its use is highly discouraged.
End user licence agreement in preparation...
Feedback
Find a bug, a typo? Have a feature request? Contact us or
Create an issue on our bug tracker